HKU Chemists Unlock the Secret to Designing Ultra-Tough and Responsive “Smart” Materials"
From household plastic packaging to the flexible frameworks that support wearable electronics, polymer materials form the invisible backbone of modern life. At a microscopic level, polymers consist of long, ribbon-like molecular chains that are entangled into a disorganised mass resembling a bowl of cooked noodles. For decades, these unpredictable molecular twists and knots have made it difficult for scientists to control, map, or customise the behaviour of the final material. A research team led by Professor Yufeng WANG and Professor Ho Yu AU-YEUNG from the Department of Chemistry at The University of Hong Kong (HKU) has achieved a breakthrough to address this challenge. By using discrete molecular rings as precise structural models of polymer knots, the team untangled the complex relationship between molecular architecture and material properties, allowing them to correlate characteristics such as stiffness, strength, and elasticity with the specific structures and topologies of the molecular rings. Their findings were recently published in the prestigious Journal of the American Chemical Society (JACS). Tuning Materials with a “Metal Switch” At the heart of this research is the discovery that the “hidden length” of the rings, a form of molecular slack within the material’s structure that releases when pulled under force. Much like a seatbelt catching to absorb an impact or a spring snapping back into place, different molecular architectures respond to mechanical stress in very different ways, thereby altering how the final material behaves. By replacing the unstructured tangles in conventional polymers with molecular rings of precise structures, the researchers were able to observe how different architectures store and release energy. Simple macrocyclic rings, for instance, are highly flexible and harbour significant hidden length; when the material is subjected to stress, this internal slack unfurls to absorb the impact, resulting in exceptional toughness and durability. In contrast, mechanically interlocked rings, known as catenanes, adopt a much more constrained and compact configuration. The team found that because these interlocked rings have less “slack” to unfurl, they behave like rapid-response springs. This creates a material with high elasticity, allowing it to snap back efficiently to its original shape after being stretched. The team took the research a step further by demonstrating that these materials can be tuned on demand. By introducing copper ions to the molecular rings, the internal slack can be effectively locked in place to increase rigidity. This ability to manipulate structural rigidity enables a material’s properties—such as stiffness and elasticity—to be dynamically altered in a controllable, responsive manner. Paving the Way for Soft Robotics and Tissue Engineering This discovery provides a blueprint for creating a new generation of “smart” materials with highly specialised functions. By identifying these distinct mechanical pathways, the HKU team has provided a new framework for guiding the design of new materials with specific properties. Professor Ho Yu Au-Yeung from the HKU Department of Chemistry said the research helps scientists gain a deeper understanding of how entanglements at the molecular level influence material properties, opening up new possibilities for designing materials with specialised functions for different applications. “By choosing the right molecular ‘knots’ and controlling their ‘hidden length’, we are now able to design materials with specific functions tailored to different needs.” Professor Yufeng Wang from the HKU Department of Chemistry added that the findings could have important implications for fields such as soft robotics, tissue engineering and wearable electronics. “For example, soft robots require materials that are both flexible and strong; tissue engineering materials need to mimic the complex and dynamic movements of human muscles; while wearable electronic devices require both high durability and elasticity. This research provides scientists with a new framework for designing smart materials with specialised functions for different applications.” The research work is supported by a Collaborative Research Fund (C7075-21G) and General Research Fund (17313222) of the Research Grants Council of Hong Kong and HKU-CAS Joint Laboratory on New Materials (JLFS/P-701/24). First authors of the paper (Tianjin LUO, Yulin DENG and Mingda HU) are PhD students at the HKU Department of Chemistry. For more details, please refer to the journal paper “Role of Molecular Topology Elucidated in Unified Gels” published in the Journal of the American Chemical Society.
NEWS DETAIL

HKU Science Joins China’s First Mars Sample Return Mission Tianwen-3
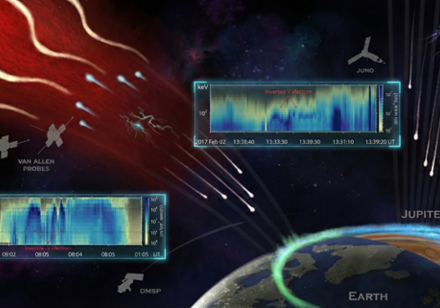
According to the selection results recently released by the China National Space Administration, HKU Science will participate in China’s first Mars Sample Return mission, Tianwen-3. The Short-Wavelength Infrared Spectrometer, led and developed by the HKU Department of Earth and Planetary Sciences (DEPS), has been selected as a payload for deployment on the service module of the Tianwen-3 mission. The orbital spectrometer will monitor Martian dust storms to support safe landing operations, conduct high-resolution mineralogical mapping of candidate landing sites, and continue long-term observations of Mars’s low-latitude regions after the sample return phase. It will play a critical role in searching for biosignatures, detecting hydrous minerals, and surveying Martian resources. The project is led by Professor Yiliang LI of the Department of Earth and Planetary Sciences at HKU, with major collaborating institutions including Zhejiang University and the Chinese Academy of Sciences’ Changchun Institute of Optics, Fine Mechanics, and Physics. In addition, the Tianwen-3 orbiter will carry three collaborative payloads, including the COSPAR-led Mars PEX Spectrometer, in which the HKU Laboratory for Space Research, led by Professor Quentin Parker, participates alongside Shenzhen University. Designed to search for traces of life on Mars and analyse surface mineral composition, the instrument will help investigate potential organic compounds and their distribution on the Martian surface.
NEWS DETAIL