29 Feb 2024
Cracking Epigenetic Inheritance: HKU Biologists Discovered the Secrets of How Gene Traits are Passed on

HKU biologists unlock the secrets of epigenetic inheritance. The key members of the research team at HKU include Professor Yuanliang Zhai (left) and PhD student Yuan Gao (right).
A research team led by Professor Yuanliang ZHAI at the School of Biological Sciences, The University of Hong Kong (HKU) collaborating with Professor Ning GAO and Professor Qing LI from Peking University (PKU), as well as Professor Bik-Kwoon TYE from Cornell University, has recently made a significant breakthrough in understanding how the DNA copying machine helps pass on epigenetic information to maintain gene traits at each cell division. Understanding how this coupled mechanism could lead to new treatments for cancer and other epigenetic diseases by targeting specific changes in gene activity. Their findings have recently been published in Nature.
Background of the Research
Our bodies are composed of many differentiated cell types. Genetic information is stored within our DNA which serves as a blueprint guiding the functions and development of our cells. However, not all parts of our DNA are active at all times. In fact, every cell type in our body contains the same DNA, but only specific portions are active, leading to distinct cellular functions. For example, identical twins share nearly identical genetic material but exhibit variations in physical characteristics, behaviours and disease susceptibility due to the influence of epigenetics. Epigenetics functions as a set of molecular switches that can turn genes on or off without altering the DNA sequence. These switches are influenced by various environmental factors, such as nutrition, stress, lifestyle, and environmental exposures.
In our cells, DNA is organised into chromatin. The nucleosome forms a fundamental repeating unit of chromatin. Each nucleosome consists of approximately 147 base pairs of DNA wrapped around a histone octamer which is composed of two H2A-H2B dimers and one H3-H4 tetramer. During DNA replication, parental nucleosomes carrying the epigenetic tags, also known as histone modifications, are dismantled and recycled, ensuring the accurate transfer of epigenetic information to new cells during cell division. Errors in this process can alter the epigenetic landscape, gene expression and cell identity, with potential implications for cancer and ageing. Despite extensive research, the molecular mechanism by which epigenetic information is passed down through the DNA copying machine, called the replisome, remains unclear. This knowledge gap is primarily due to the absence of detailed structures that capture the replisome in action when transferring parental histones with epigenetic tags. Studying the process is challenging because of the fast-paced nature of chromatin replication, as it involves rapid disruption and restoration of nucleosomes to keep up with the swift DNA synthesis.
In previous studies, the research team made significant progress in understanding the DNA copying mechanism, including determining the structures of various replication complexes. These findings laid a solid foundation for the current research on the dynamic process of chromatin duplication.
Summary of Research Findings
This time, the team achieved another breakthrough by successfully capturing a key snapshot of parental histone transfer at the replication fork. They purified endogenous replisome complexes from early-S-phase yeast cells on a large scale and utilised cryo-electron microscopy (cryo-EM) for visualisation.
They found that a chaperone complex FACT (consisting of Spt16 and Pob3) interacts with parental histones at the front of the replisome during the replication process. Notably, they observed that Spt16, a component of FACT, captures the histones that have been completely stripped off the duplex DNA from the parental nucleosome. The evicted histones are preserved as a hexamer, with one H2A-H2B dimer missing. Another protein that involved in DNA replication, Mcm2, takes the place of the missing H2A-H2B dimer on the vacant site of the parental histones, placing the FACT-histone complex onto the front bumper of the replisome engine, called Tof1. This strategic positioning of histone hexamer on Tof1 by Mcm2 facilitates the subsequent transfer of parental histones to the newly synthesised DNA strands. These findings provide crucial insights into the mechanism that regulates parental histone recycling by the replisome to ensure the faithful propagation of epigenetic information at each cell division.
This study, led by Professor Zhai, involved a collaborative effort that spanned nearly eight years, starting at HKUST and concluding at HKU. He expressed his excitement about the findings, ‘It only took us less than four months from submission to Nature magazine to the acceptance of our manuscript. The results are incredibly beautiful. Our cryo-EM structures offer the first visual glimpse into how the DNA copying machine and FACT collaborate to transfer parental histone at the replication fork during DNA replication. This knowledge is crucial for elucidating how epigenetic information is faithfully maintained and passed on to subsequent generations. But, there is still much to learn. As we venture into uncharted territory, each new development in this field will represent a big step forward for the study of epigenetic inheritance.’
The implications of this research extend beyond understanding epigenetic inheritance. Scientists can now explore gene expression regulation, development, and disease with greater depth. Moreover, this breakthrough opens up possibilities for targeted therapeutic interventions and innovative strategies to modulate epigenetic modifications for cancer treatment. As the scientific community delves deeper into the world of epigenetics, this study represents a major step towards unravelling the complexities of replication-coupled histone recycling.
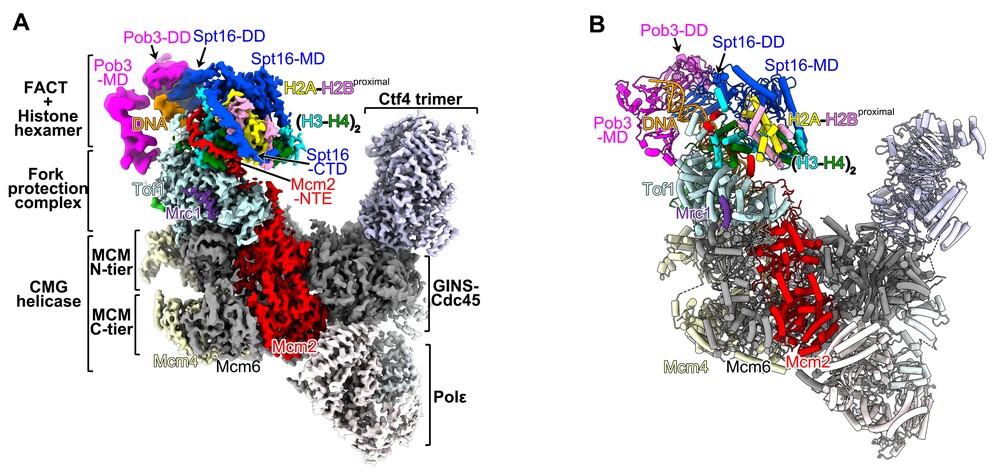
The cryo-EM structure of the yeast replisome in complex with FACT and parental histones (A) and its atomic model (B). Modified from Li et al, Nature (2024)
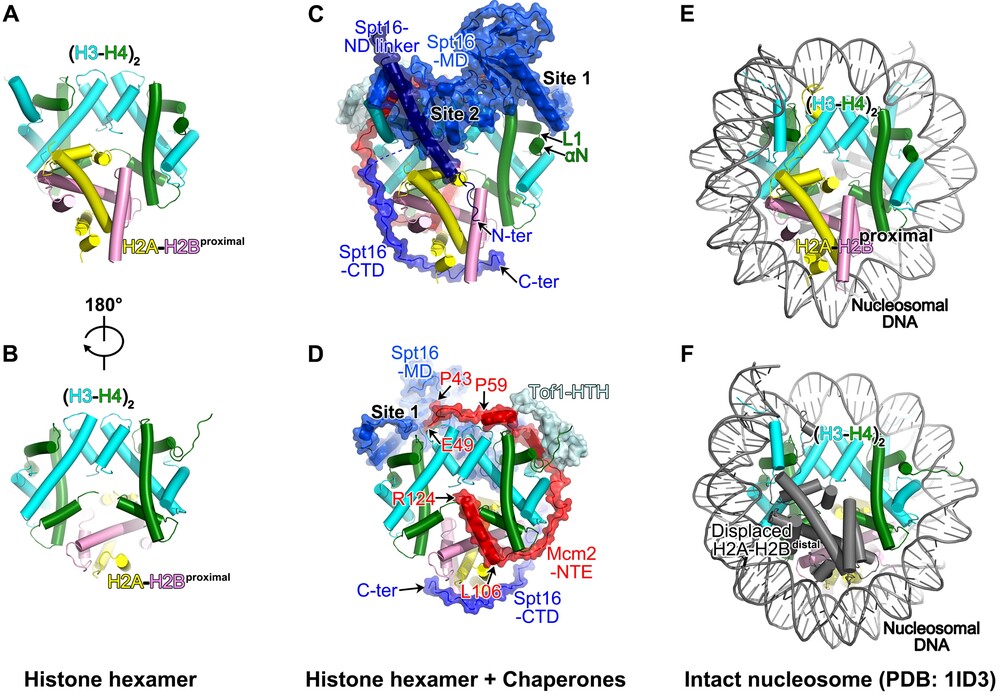
The evicted histone hexamer and its chaperons from the replisome structure. (A-B) The architecture of the parental histone hexamer. (C-D) The histone-chaperone complex on the replisome. (E-F) The structure of an intact nucleosome. Modified from Li et al, Nature (2024)
About the Research Team
Apart from Professor Yuanliang Zhai’s lab, the research team also includes Professor Xiang David Li from Department of Chemistry of HKU, Professor Yang Liu and Professor Keda Zhou from School of Biomedical Sciences of HKU, Professor Shangyu Dang from Division of Life Science of HKUST, and others. Learn more about Professor Yuanliang Zhai’s work and his research team:
https://www.scifac.hku.hk/people/zhai-yuanliang or https://zhai95.wixsite.com/mysite-1
Co-authors include Mr Yuan Gao, Mr Jian Li, Dr Zhichun Xu from School of Biological Sciences (SBS) of HKU; Dr Ningning Li, Ms Yujie Zhang, Dr Jianxun Feng from School of Life Sciences of PKU, Dr Daqi Yu and Dr Jianwei Lin from Department of Chemistry of HKU, and Dr Yingyi ZHANG from Biological Cryo- EM Center of HKUST.
The journal paper can be accessed here.