30 Jan 2023
Catching the wrongdoers in the act: HKU chemists develop a novel tool to decipher bacterial infections in real time
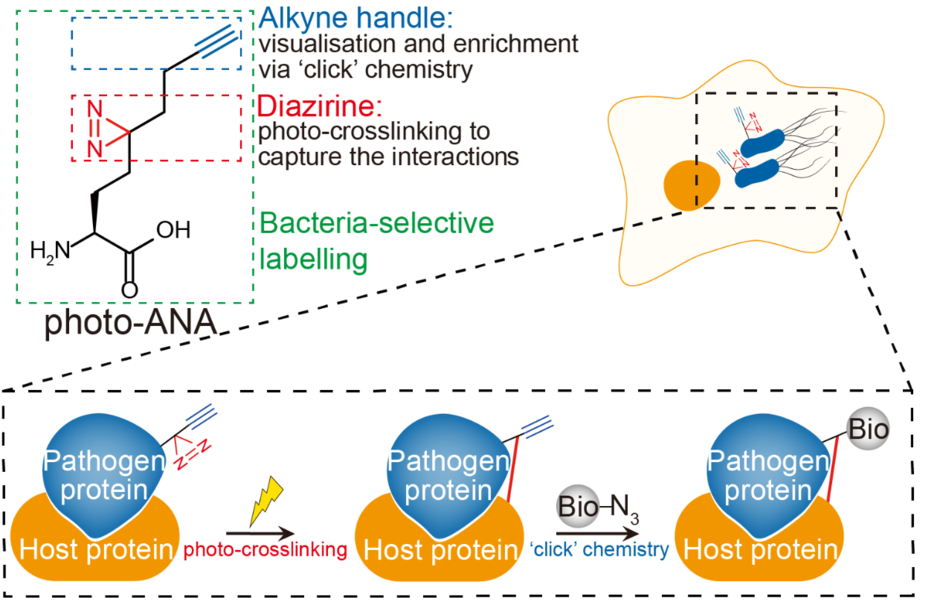
Image modified from original illustration of Li et al, 2023 Nature Chemical Biology (doi: 10.1038/s41589-022-01245-7)
A research team led by Professor Xiang David LI from the Department of Chemistry at The University of Hong Kong (HKU) has developed a novel chemical tool to reveal how bacteria adapt to the host environment and control host cells. This tool can be used to investigate bacterial interactions with the host in real-time during an infection, which cannot be easily achieved by other methods. The findings were recently published in a top-class international scientific journal — Nature Chemical Biology.
Pathogenic bacteria, while only numbering fewer than a hundred, heavily threaten human health all over the world. For example, Mycobacterium tuberculosis infection causes tuberculosis, which leads to more than one million deaths each year. It was the world’s deadliest infectious disease until it was overtaken by COVID-19. Despite effective antibiotic treatments, multiple drug-resistant tuberculosis has become a growing problem worldwide. Therefore, a more comprehensive understanding of how bacteria infect their human host is key to developing new drugs and therapies.
When bacteria meet their host (e.g., human cells), they send out ‘assassins’ (virulence factor proteins) that ‘hijack’ important protein players of the host to sow chaos during an invasion. Therefore, investigating which virulence factors bacteria secrete and which host proteins are targeted is crucial for the understanding of bacterial infections. However, it can be extremely challenging to identify these key players among the ‘crowded streets’ (excessive host cellular matrix).
To tackle this challenge, Professor Li’s group designed a multifunctional unnatural amino acid called photo-ANA that only labels proteins of the engineered bacteria but not the host during infection. With the help of its alkyne handle, photo-ANA can conjugate with fluorescence or biotin via a Nobel prize-winning chemical reaction (‘click’ chemistry), which enables the visualisation and enrichment of the labelled bacterial proteins from the complex host environment. Thus, Photo-ANA serves as an ‘undercover agent’ to gather intelligence and tag all the ‘assassins’ sent by the bacteria. More importantly, photo-ANA also carries a diazirine group that can ‘handcuff’ the bacterial virulence proteins to their host target proteins upon exposure to ultraviolet (UV) light — cathing them in the act.
Using photo-ANA, Professor Li’s group comprehensively profiled the adaptation of Salmonella, bacteria that can cause severe diarrhoea, to the host environment and revealed the extensive interplay between Salmonella and the host during different infection stages, which identified known interactions and some newly discovered interactions. Moreover, the photo-ANA-based approach can be easily applied to other pathogenic bacteria and even other pathogens such as fungi.
With this new chemical tool, scientists can now investigate the activity of bacteria inside the host in real-time. In the future, this tool may help us decipher the hidden interactions of deadly bacteria with the host and the mechanisms of multidrug-resistant superbugs, which will further our understanding of infectious diseases and inspire new treatments.
Link of journal paper can be accessed from here.







